Register (Free Test Slot)
Startup Readiness And Recommendations
New slots need time before DP recommendations become meaningful. A prediction episode is a forecast that later receives real CGM values as the answer. A +120 minute episode cannot finish until about two hours after the forecast was made, so runtime MAPE, bias, therapy suggestions and recommendation gates lag behind live BG.
| Stage | Expected | If Missing |
|---|---|---|
| Right after setup | Latest BG, profile ISF/CR/basal, AAPS/Loop IOB/COB if devicestatus is fresh. | Check Nightscout URL/token, DNS, CGM age and data_aaps_age_seconds. |
| After first train | DP model forecast plus model_val_mae and model_val_p95. |
Run train and check that Nightscout has enough CGM history. |
| After 2-6 hours | First finalized episodes, prediction_mape and forecast error metrics. |
Wait for future CGM to arrive; each episode needs enough actual points. |
| After about 12 episodes | ISF/basal suggestions can start. CR suggestions need enough carb entries. | Check BIAS_MIN_EPISODES_TOTAL, BIAS_GLOBAL_FALLBACK and carb treatments. |
| After 20-30 episodes | Recommendation gate can unlock if +60m MAPE is acceptable. | Check recommendation_gate_episodes, prediction_mape{60} and gate_reason. |
| After 3-14 days | Suggestions become more useful across different hours, meals and activity levels. | Sparse hours still use profile values or global bias until more data exists. |
Recommendation Gate
Start conservative: RECOMMENDATION_GATE_MIN_EPISODES=20 and
RECOMMENDATION_GATE_MAX_MAPE60=0.30. Lower episodes only for testing. Higher MAPE allows
recommendations from a less accurate model.
Therapy Suggestions
BIAS_MIN_EPISODES_TOTAL=12 allows early bias learning. Per-hour bias needs
BIAS_MIN_EPISODES_PER_HOUR=3. With BIAS_GLOBAL_FALLBACK=true, sparse hours use
overall bias until enough hourly data exists.
Carb Ratio
CR_SUGGEST_MIN_CARB_EVENTS=5 protects CR changes. If carbs are not entered or not published
to Nightscout, CR suggestions should stay close to the profile.
Common Block Reasons
episodes<N means wait for more finalized episodes. uam_detected blocks when BG
rises without enough carbs/COB. degenerate_forecast means the raw model output is unhealthy.
Stale CGM must be fixed before trusting output.
Grafana Examples
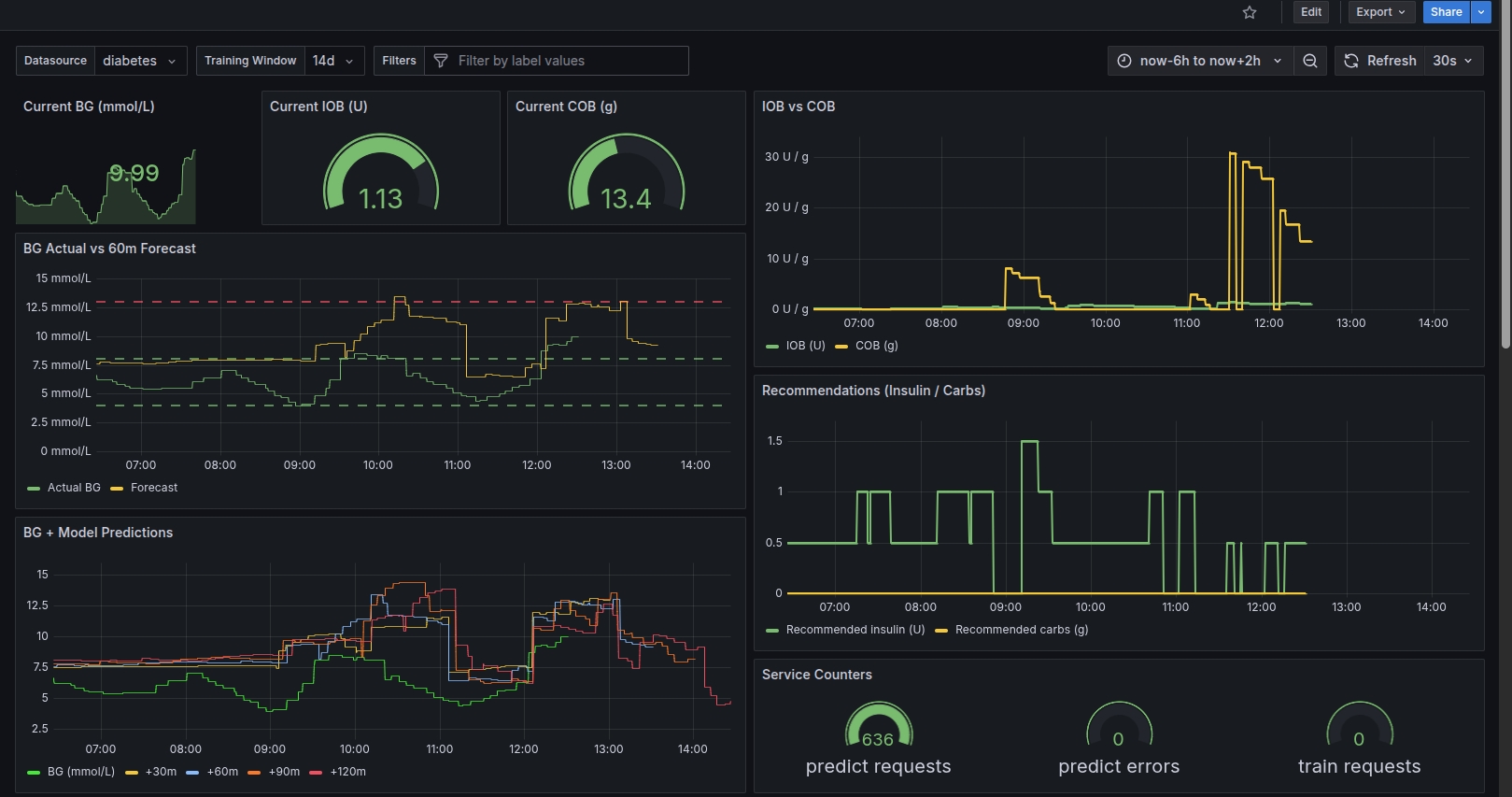
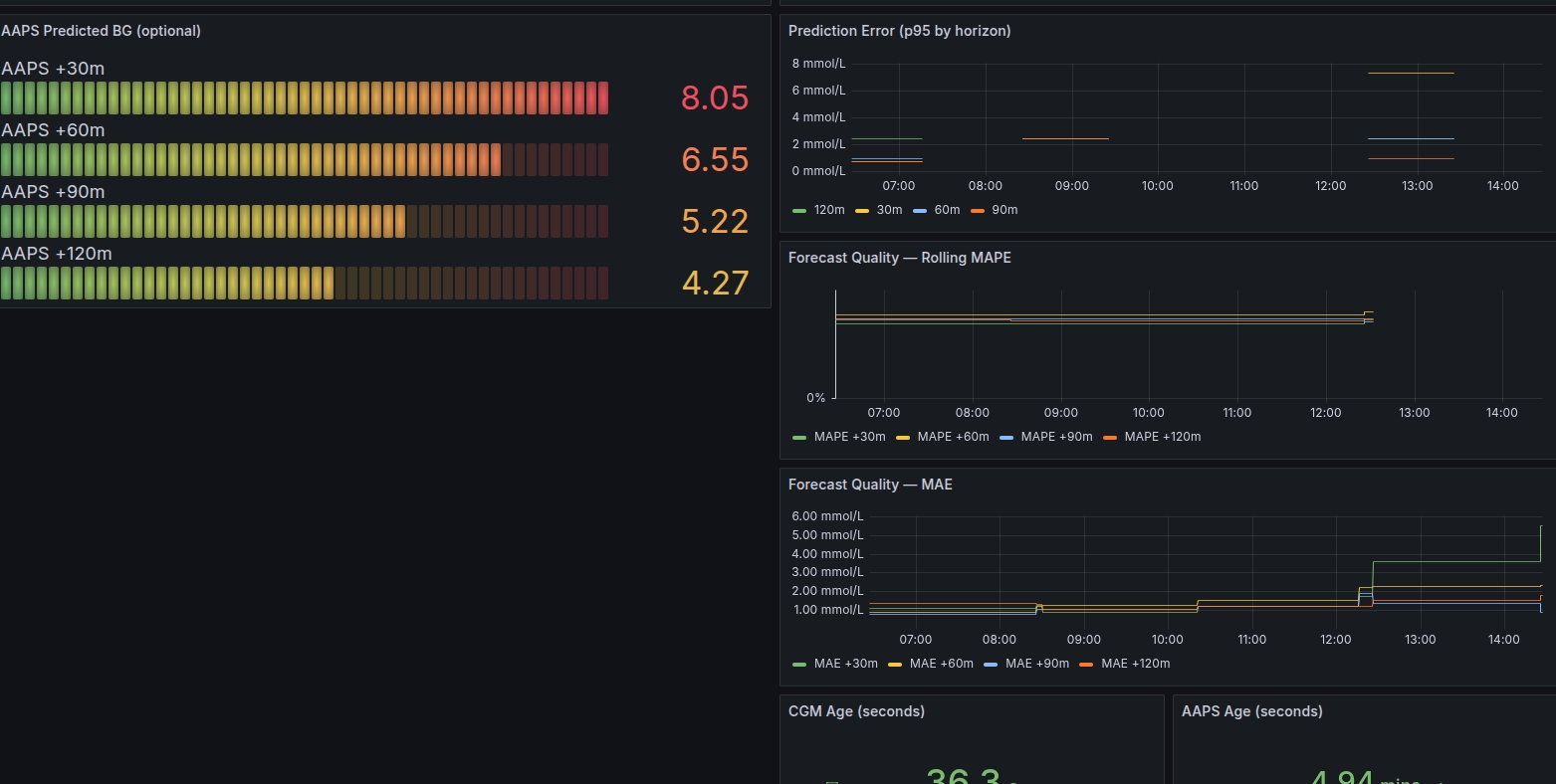
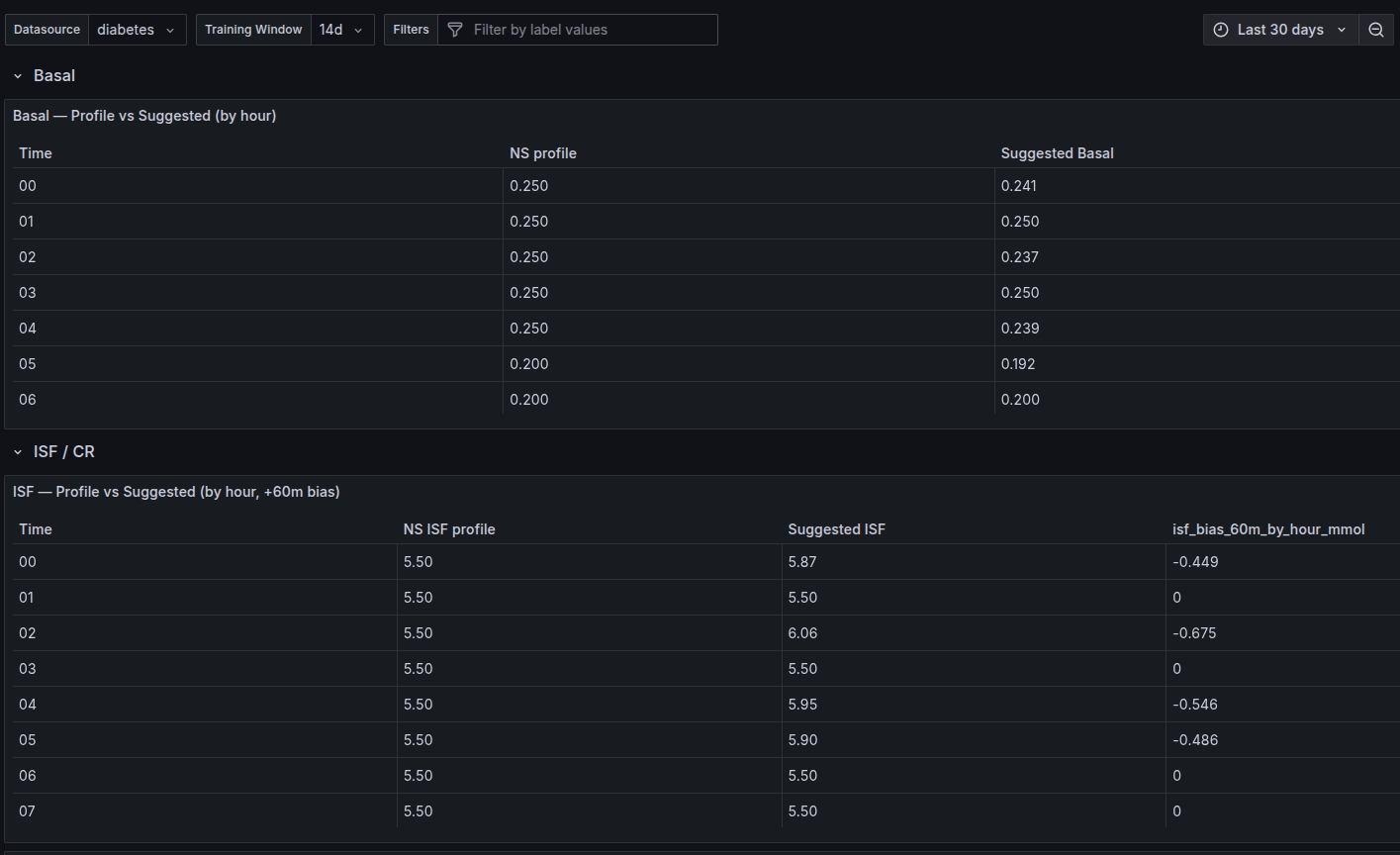
How It Works (Technical)
- Nightscout is the source of truth (SGVs, treatments, profile, devicestatus).
- We resample to 5‑minute steps and build features: BG, IOB/COB, SMB/bolus, carbs, ISF/CR, basal, targets, time + trend.
- The active runtime model is an LSTM + Attention sequence model. GRU, TCN/Conv1D and ensemble models are planned challenger models.
- Predictions are checked against actual CGM outcomes with MAE, MAPE, p95 error, hourly bias and fallback/health metrics.
- Therapy hints are quality-gated and stay as decision support; DP is not a medical device or automated dosing system.
Training
/train fetches recent Nightscout history, merges CGM and treatments, adds profile and trend
features, builds sliding windows from LOOKBACK, trains with MAE loss, validates on the newest
data, then saves latest.keras plus metadata.
Prediction
/predict builds the latest feature window, prefers fresh AAPS/Loop IOB/COB and targets when
available, runs the LSTM forecast, applies learned bias correction, and falls back to cached/AAPS/flat
forecasts if the model output is unhealthy.
Quality Metrics
Grafana/Prometheus track model_val_mae, model_val_p95,
forecast_error, prediction_mape, +60m hourly bias,
stale CGM/AAPS age, floor hits, NaNs, constant input columns and fallback usage.
Therapy Suggestions
Bias patterns drive bounded suggested ISF, carb ratio and basal schedules. Recommendation output is blocked when CGM is stale, the forecast is degenerate, there are too few completed episodes, or +60m MAPE is above the configured gate threshold. UAM detection can also block recommendations when BG rises without recent carbs or COB.
Outcome Roadmap
Next best upgrades are champion/challenger deployment, a GRU challenger, a TCN/Conv1D challenger, AAPS/model/trend ensemble selection, uncertainty scoring and weighted training around meals, boluses, lows, highs and rapid BG movement.
Model Inputs
Core inputs include bg, iob, cob, smb,
bolus, carbs, isf, carb_ratio,
basal, target, time-of-day, weekday, time since meal/bolus and BG deltas over
5, 15 and 30 minutes.
Full Technical Solution
Current Runtime Model
The active predictor is EnhancedBGPredictor in app/models.py. Runtime currently
uses an LSTM + Attention sequence model. GRU is not active today; it is a planned challenger model.
Input: LOOKBACK rows x feature_count default LOOKBACK=864 = about 72 hours at 5-minute cadence Model: LSTM(128, return_sequences=True) BatchNormalization Dropout(0.3) LSTM(64, return_sequences=True) LSTM(32) query branch LSTM(32) value branch Attention LSTM(64) Dense(PREDICTION_HORIZON_STEPS) Output: one predicted BG value per future 5-minute step default PREDICTION_HORIZON_STEPS=18 = 90 minutes 24 steps = 120 minutes
Training Pipeline
Training starts at /train, calls train_from_nightscout(), fetches Nightscout
CGM/treatments/profile, normalizes BG to mmol/L, builds feature rows, creates sliding windows and trains
with MAE loss.
- Fetch CGM entries for
TRAINING_WINDOW_DAYS. - Fetch treatments and the active Nightscout profile.
- Normalize timestamps to UTC, sort and de-duplicate CGM rows.
- Merge treatments into the CGM timeline and build IOB, COB, SMB, bolus, carbs and basal columns.
- Add profile, time and BG trend features.
- Build
Xas pastLOOKBACKrows andyas future BG values. - Validate on newest data using
VALIDATION_DAYSorVALIDATION_SPLIT. - Save timestamped Keras model, update
models/latest.kerasand write model metadata JSON.
Model Features
bg, iob, cob, smb, bolus, carbs, isf, carb_ratio, basal, target, hour_sin, hour_cos, wday_sin, wday_cos, time_since_last_meal, time_since_last_bolus, d_bg_5m, d_bg_15m, d_bg_30m, current_bg
The most outcome-critical inputs are usually BG trend, IOB, COB, active target, recent bolus/carb timing, active ISF/CR/basal profile and whether AAPS/Loop devicestatus is fresh.
Prediction Pipeline
/predictgets recent CGM, treatments over the DIA window and the active profile.- Fallback IOB/COB is calculated from treatments.
- Fresh AAPS/Loop devicestatus can override IOB/COB and provide targets, autosens, dynamic ISF and predBG.
- The latest feature window is passed into the LSTM model.
- Bias correction is applied when enough completed prediction episodes exist.
- Degenerate/floor forecasts are detected and can fall back to cached last-good, AAPS or flat forecast.
- Prometheus metrics are updated for live BG, IOB/COB, forecasts, quality and fallbacks.
- A prediction episode is stored so future actual CGM can be compared against the forecast.
Outcome Metrics
Training quality and live outcome are measured separately. Training-time metrics tell if a new model looked good during validation; runtime metrics tell if predictions matched real future CGM.
Training:
model_val_mae{horizon_min="30|60|90|120"}
model_val_p95{horizon_min="30|60|90|120"}
Runtime:
forecast_error{horizon_min="30|60|90|120"}
prediction_mape{horizon_min="30|60|90|120"}
recommendation_gate_mape_60
recommendation_gate_episodes
isf_bias_60m_by_hour_mmol{hour="00".."23"}
prediction_bias_applied{horizon_min="30|60|90|120"}
UAM / Unannounced Meal Detection
DP detects a first-pass UAM pattern at runtime without changing the trained model shape. It computes a weighted UAM score from BG rise, AAPS/Loop predicted rise when available, low COB and missing recent carb entries. When enabled, UAM can block insulin/carb recommendations while still returning the forecast.
UAM_DETECTION_ENABLE=true UAM_BLOCK_RECOMMENDATIONS=true UAM_MAX_COB_G=5 UAM_RECENT_CARBS_WINDOW_MIN=90 UAM_DBG_15M_MIN_MMOL=0.6 UAM_DBG_30M_MIN_MMOL=1.0 UAM_SCORE_THRESHOLD=0.70 Metrics: uam_detected uam_score uam_rise_rate_15m_mmol uam_rise_rate_30m_mmol uam_recent_carbs_grams uam_minutes_since_last_carbs
MAE, MAPE, p95 and Bias
- MAE: average absolute error in mmol/L.
- MAPE: error relative to actual BG.
- p95: how bad the worst 5 percent of predictions are.
- Bias: whether actual BG is systematically higher or lower than predicted by hour.
Bias is calculated as actual - predicted at +60 minutes. Positive bias means DP under-predicted;
negative bias means DP over-predicted. This drives bounded suggested ISF, carb ratio and basal schedule
changes.
Safety and Recommendation Gate
Therapy hints are blocked when quality is not good enough. The gate can require enough completed episodes, acceptable +60m MAPE, fresh CGM and a non-degenerate forecast.
RECOMMENDATION_GATE_ENABLE=true RECOMMENDATION_GATE_WINDOW_DAYS=7 RECOMMENDATION_GATE_MIN_EPISODES=30 RECOMMENDATION_GATE_MAX_MAPE60=0.30
Prediction Health Metrics
prediction_floor_hits_60min prediction_degenerate prediction_using_cached prediction_using_aaps_fallback prediction_using_flat_fallback prediction_input_nan_cells prediction_input_constant_cols data_cgm_age_seconds data_aaps_age_seconds data_cgm_missing_total data_treatments_skipped_total
Best Next Improvements
- Champion/challenger model selection before replacing a working model.
- Add GRU as a real challenger with
MODEL_ARCH=lstm|gru. - Add TCN/Conv1D/WaveNet-style challenger for fast time-series inference.
- Compare LSTM, AAPS/Loop forecast, flat current-BG and trend baseline in an ensemble selector.
- Add prediction uncertainty/confidence and use it to harden the recommendation gate.
- Weight meals, boluses, SMB, lows, highs and rapid BG movement more heavily during training.
LLMs are useful for explanation and reporting, but not as the numeric BG forecast model. The prediction path should stay in measurable time-series logic with MAE, MAPE, p95, bias and explicit safety gates.
Build Notes
- API (Flask): /predict + /train.
- Poller runs the predict → report loop and logs actual outcomes.
- Prometheus stores metrics (BG, IOB/COB, forecasts, errors).
- Grafana dashboards show predictions + therapy suggestions.
- When editing settings: use
NIGHTSCOUT_URLwithout trailing/(example:https://site.com).
